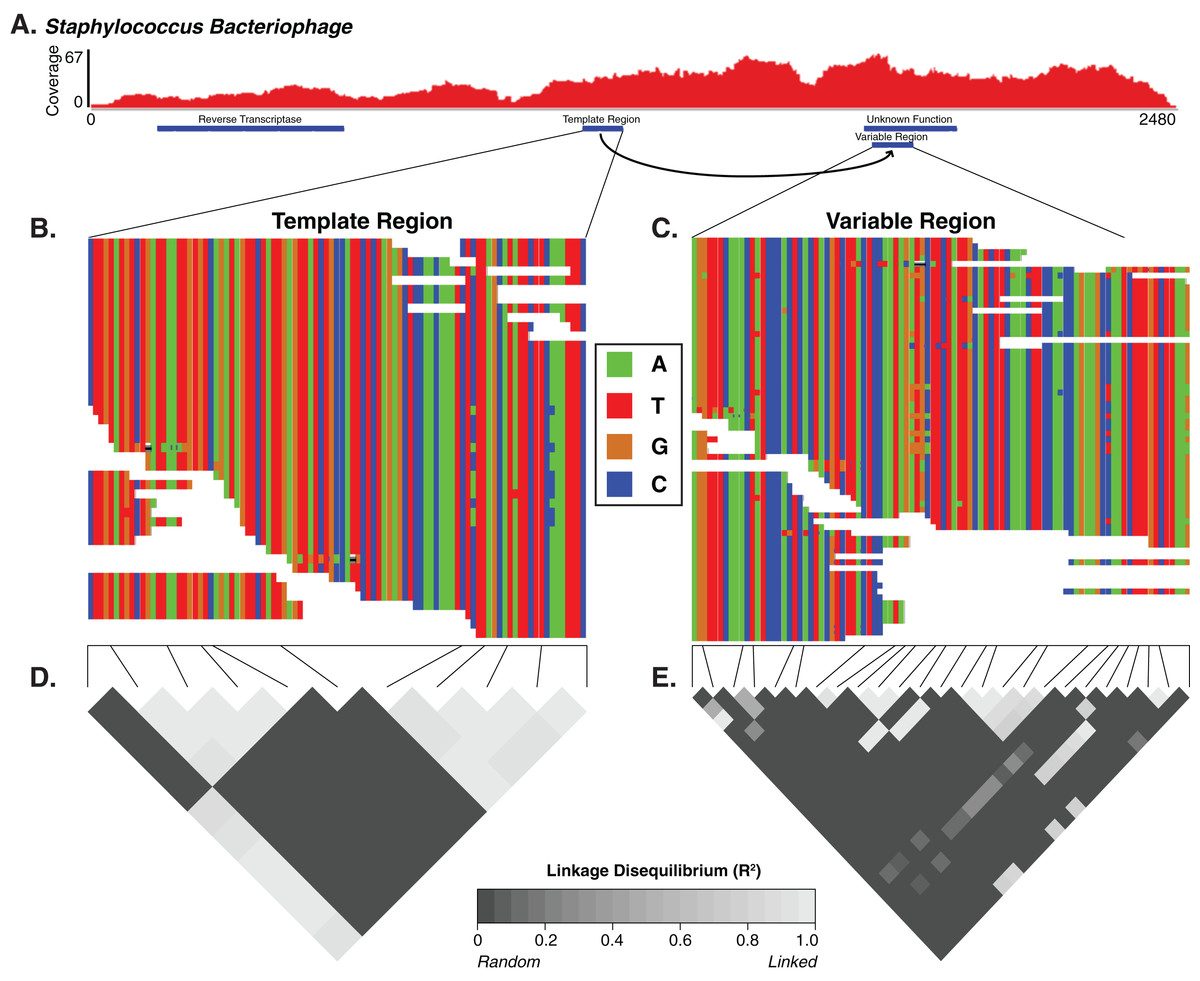
 |
| We identified diversity generating retroelements as a potential mechanism driving targeted genomic diversity. |
A few weeks ago some colleagues and myself published a new manuscript looking at the diversity of the human skin virome. In our previous previous work, we evaluated the diversity of viruses on the skin. Other groups have looked at virus diversity at other body sites including the gut, lungs, and oral cavity. Our new paper focused on the diversity within viruses on the skin. It provided initial insight into the genomic variability associated with major viruses in the skin virome. In other words, it was a "high resolution" study of the virome.
One of the highlights of the manuscript was identifying numerous hyper-variable loci within the skin virus genomes that we investigated. We did this using a SNP geometric distribution approach instead of a sliding window because it allowed us to establish regions that were more variable than would be expected by random chance, and did not require us to arbitrarily establish a window size for the loci. The loci identified using this approach were associated with stronger evolutionary pressure than their adjacent regions, suggesting they are functionally important. We followed up with this, but I will let you get the details from the manuscript.
A methodological highlight was the validation of our findings using an existing dataset from a different lab. We performed our analytical workflow a second time using another skin metagenomic dataset from a different skin microbiome lab. Even though the second dataset did not undergo virus purification, we were still able to pull out enough viral reads to perform our targeted analysis. We replicated our findings in the second dataset, thereby supporting the strength and biological ubiquity of our findings. My hope is that more groups will perform this type of validation in their future studies, especially since there is so much archived data just waiting to be utilized.
The challenge with this study was writing the analytical tools that I needed to answer our questions. In the end, I built a lot of tools that allowed me to answer evolutionary and functional virome questions. I think these are pretty easy to use, and made them freely available on GitHub. If you are interested in performing similar evolutionary analyses on your virome datasets, check the code out here and let me know if you have any questions.
In the end, I think this is a pretty cool study and I really enjoyed working on it. If this summary sounds interesting, I suggest you check out the paper. It is freely available online and easy to download.
As always, if you have any questions, comments, or concerns, please let me know in the comments section below, shoot me an email, or find me on twitter (links are to the right). I always love to hear from readers!
References
Hannigan GD, Zheng Q, Meisel JS, Minot SS, Bushman FD, & Grice EA (2017). Evolutionary and functional implications of hypervariable loci within the skin virome. PeerJ, 5 PMID: 28194314
This is the most skin cell infecting virus and it's important to take care of your skin at early ages. Please guide a cure for this virus. Thanks
ReplyDeleteHi Friends i am so glad to writing this article today to tell the world how Dr voodoo cured my HSV VIRUS,i have been detected with HSV-1 AND HSV-2 since five years ago, ever since then my life has been in complete bizarre and agony,i have used so many drugs that was prescribed to me by several doctors,but it didn't cure my HSV VIRUS neither did it reduce the pain,until a certain i was checking for solution in the internet,then miraculously came across Dr voodoo the powerful herbalist that cure numerous individuals HSV-1 AND HSV-2 INFECTION,then i contacted his whatsApp number at +2348140120719 or email: voodoospelltemple66@gmail.com i explained everything to him and prepared a cure that cure my HSV-1 AND HSV-2 disease totally after receiving his herbal medicine, so my friends viewers why wait and be suffer when there is someone like Dr voodoo that can cure any disease HIV/ CANCER/ HEPATITIS B VIRUS, you can contact his via : voodoospelltemple66@gmail.com or WHATSAPP +2348140120719
DeleteHERPES INFECTION CURE IS 100% AVAILABLE WITH THE HELP OF DR OGBERAESE HERBAL MEDICINE.Joy and happiness is all i can see around ever since i came in contact with this great man Called Dr Ogberaese. i complained bitterly to him about me having herpes only for him to tell me it’s a minor stuff. He told me he has cured thousands of people but i did not believe until he sent me the herbal medicine and i took it as instructed by this great man, only to go to the hospital after two weeks for another test and i was confirmed negative. For the first time in four years i was getting that result. i want to use this medium to thank this great man. His name is Dr Ogberaese , i came in with his email through a friend in USA called Edith Rose and ever since then my live has been full with laughter and great peace of mind. i urge you all with herpes or HSV1&2 to contact him if you are willing to give him a chance. you can contact him through his email drogberaese54@gmail.com or you can also WhatsApp him +2348073818653
DeletePowerful Herbal treatment is 100% guarantee for HSV cure, the reason why most people are finding it difficult to cure HSV 1 an 2 is because they believe on medical report, drugs and medical treatments which is not helpful to cure HSV and hasn't proved any sign of helping. Natural roots/herbs are the best remedy which can easily eradicate herpes forever. I never believed it until I was helped and cured of my 16 months genital herpes with natural herbal medicines from Dr oduku. Where other medical prescribed drugs and treatments failed, Dr oduku natural herbs helped saved me from Genital herpes permanently and I’m so grateful for this. You can also get help from this great and powerful Herbalist Dr oduku by reaching him on whatsApp +2348124765093 also EMAIL: drodukuherbalhealingcentre@gmail.com or visit his page https://www.facebook.com/Droduku/ or his tiktok.com/@drodukuhome
DeleteDr Oduku is the best herbal doctor.
Here is my private e-mail laurielkeene159@gmail.com you can also get in touch with me.
call +2348038253815 or add us on whatsApp +2348038253815 or email illuminaticult0666@gmail.com GREETINGS!!!!! FROM THE GREAT GRAND MASTER! IN REGARDS OF YOU BECOMING A MEMBER OF THE GREAT ILLUMINATI, WE WELCOME YOU. Be part of something profitable and special (WELCOME TO THE WORLD OF THE ILLUMINATI). Are you a POLITICIAN, ENGINEER,DOCTOR, ENTERTAINER,MODEL,GRADUATE/ STUDENT,OR YOU HAVE IT IN MIND TO EXPAND YOUR BUSINESS/COMPANIES TO BECOME GREAT MINDS. It is pertinent to also know that For becoming a member, you earn the sum of $1,000,000 as the illuminati membership salary monthly.Be a part of this GOLDEN “OPPORTUNITY” The great illuminati Organization makes you rich and famous in the world, it will puxll you out from the grass root and take you to a greater height were you have long aspired to be and together we shall rule the world with the great and mighty power of the Illuminati, long life and prosperity here on earth with eternal life and jubilation. You can reach Us on illuminaticult0666@gmail.com
ReplyDeleteThis is another testimony on how Chief Dr Lucky cured My HIV Disease. Do you need a cure for your HIV disease? Do you want to be cured from your cancer disease? Or you want to be free from any type of disease. Kindly visit his website https://chiefdrluckyherbaltherapy.wordpress.com/ , he just cured my HIV disease and I’m very grateful to him, he is the only herbalist that can cure you.
ReplyDeleteWhatsApp number : +2348132777335
Via Email : chiefdrlucky@gmail.com
Thank you all for reading,
God bless"
HERPES INFECTION CURE IS 100% AVAILABLE WITH THE HELP OF DR OGBERAESE HERBAL MEDICINE.Joy and happiness is all i can see around ever since i came in contact with this great man Called Dr Ogberaese. i complained bitterly to him about me having herpes only for him to tell me it’s a minor stuff. He told me he has cured thousands of people but i did not believe until he sent me the herbal medicine and i took it as instructed by this great man, only to go to the hospital after two weeks for another test and i was confirmed negative. For the first time in four years i was getting that result. i want to use this medium to thank this great man. His name is Dr Ogberaese , i came in with his email through a friend in USA called Edith Rose and ever since then my live has been full with laughter and great peace of mind. i urge you all with herpes or HSV1&2 to contact him if you are willing to give him a chance. you can contact him through his email drogberaese54@gmail.com or you can also WhatsApp him +2348073818653
ReplyDelete